Increased DAPK1 but decreased CCL2 plasma levels of nucleic acids in patients with stable angina
Darko Cerne
[*]
[1]
Irma Stern
[1]
Igor Kranjec
[2]
Andreja Cerne
[2]
Darko Zorman
[2]
Simona Krzisnik-Zorman
[2]
Janja Marc
[1]
Introduction
Death-associated protein kinase (encoded by DAPK1) is a proapoptotic, calmodulin-regulated serine/threonine kinase that carries a C-terminal death domain. Apoptosis has been identified as a prominent feature of atherosclerosis (1) and DAPK1 mRNA and DAPK1 are increased in carotid atherosclerotic plaques (2). Monocyte chemotactic protein-1 (encoded by CCL2) is involved in the formation, progression, and destabilization of atherosclerotic plaques (3). A follow-up study in hypercholesterolemic rabbits showed that the increase of CCL2 mRNA in arterial wall correlated with atherosclerotic plaque development (4).
Cell-free nucleic acids are nucleic acids (DNA, mRNA) present in plasma, serum and cell-free fractions of various others biological fluids (5). As to coronary artery disease, the prognostic value of nucleic acids in theplasma of patients with acute coronary syndrome has been established (6,7) and patients with coronary atherosclerosis have increased plasma cathepsin S mRNA levels (8). We proposed that it should be possible to measure DAPK1 mRNAand CCL2 mRNA levels, i.e. from genes responsible for plaque development and destabilization, in plasma of patients with coronary atherosclerosis, and we hypothesized that these levels are increased. If the answers were positive, it would mean that plasma mRNA estimation could offer non-invasive in vivo assessment and monitoring of gene expression profile in atherosclerotic vascular beds.
Materials and methods
Patients
Materials and methods (8) and optimisation of protocols (9,10) were in detail described previously. Briefly, patients were consecutively enrolled from those with stable angina or acute coronary syndrome undergoing routine coronary angiography. Stable angina and acute coronary syndrome were diagnosed by clinical assessment, EKG evaluation and laboratory findings. Inflammatory, thyroid, liver, endocrine, neoplastic and renal diseases (serum creatinine concentration values ≥ 120 μmol/L) were excluded by detailed history and clinical examination. The study was approved by the national ethics committee. Informed consent was obtained from all participating subjects.
Methods
Peripheral blood was taken into EDTA tubes (Becton Dickinson, Plymouth, UK) after at least 12 hours of fasting and before coronary angiography. To reduce the bias resulting from instability of mRNA the same double-spin procedure was used in all subjects to obtain plasma nucleic acids. Sample processing took place within one hour of collection. Haemolysed samples were not used for analysis.
RNA was isolated from plasma usingQIAamp MinElute Virus Spin Kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations, but with the following modifications: 1) the incubation time for proteolysis was 5 min; 2) RNA was eluted from the column with 55 μL RNase-free water and the collected eluate used for a second elution of RNA from the samecolumn. Standard methods (A260/A280 on a NanoDrop or Agilent RNA 6000 Nano kit) are inappropriate for assessment of RNA yield and quality in the collected eluate because RNA carrier that is used in the process of plasma RNA isolation yields the signal, which is too dominant relative to the signal of RNA species of interest. As a consequence, RNA amounts based on starting volume rather than RNA quantity is used in all analytical procedures. Eluted mRNA was transcribed to cDNA with SuperScript VILO cDNA Synthesis Kit (Invitrogen, Carlsbad, CA, USA). First, RNA eluate was incubated in a water bath (65 ˚C) for 5 min, then immediately placed on ice for another 5 min. The manufacturer’s protocol was followed from then on.
The following TaqMan Gene Expression Assays (Applied Biosystems, Foster City, CA, USA) were used for cDNA quantification: Hs00234489_m1 for DAPK1 (exon boundary 3-4, amplicon length 105), Hs00234140_m1 for CCL2 (exon boundary 1-2, amplicon length 101) and Endogenous Control 4326316E for cyclophilin A (PPIA). cDNA was amplified by PCR according to the manufacturer’s protocol (Roche Diagnostics, GmbH, Mannheim, Germany), using a 1.5 μL (DAPK1 and PPIA) or 3.75 μL (CCL2) of the cDNA solution and the kit components in a reaction volume of 25 μL. Although the assay kits for cDNA quantification are declared to be mRNA specific, in our test conditions DNA was found to interfere in both assays. All reactions were run in triplicate and the average threshold cycle of triplicates (Ct) calculated. Between-run imprecision for the whole analytical procedure (from isolation to quantification), expressed as coefficient of variation of Ct, was less than 2%. To exclude the possibility of external nucleic acids contamination negative control (blank) was analysed in each PCR run. For clinical evaluation, the amount of each investigated transcript was standardized to the amount of a reference gene. PPIA was chosen as the most appropriate reference gene among 11 candidates available on TaqMan Human Endogenous Control Plate (Applied Biosystems), based on the smallest geometric Ct of individual candidates and results from geNorm, as in detail described previously (9).
Statistical analyses
Data are presented as counts; age is presented as median and range. Statistically significant difference between patients and controls for quantitative variables (age and plasma levels of nucleic acids) was tested using nonparametric Mann-Whitney test.
P values < 0.05 were considered statistically significant.
Statistical analyses were performed using SPSS 17.0 statistical software package (SPSS, Inc., Chicago, IL, USA).
Results
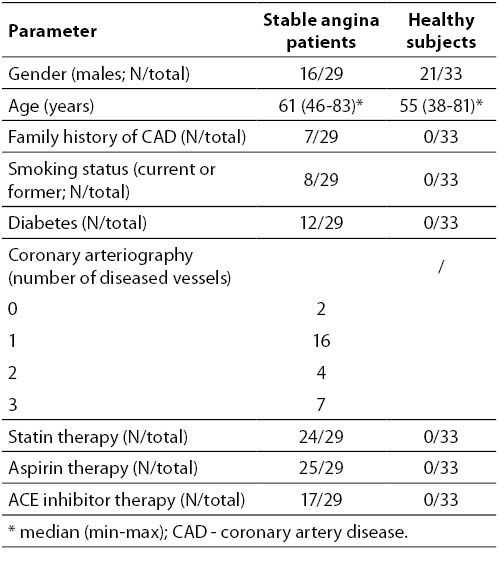
The clinical characteristics of the participants are summarized in Table 1. Healthy subjects were younger (median 55 vs. 61 years; P = 0.029). Age did not correlate with any observed parameter within either patient or control group (data not shown).
Table 1. Clinical characteristics of subjects.
Although the assay kits for cDNA quantification are declared to be mRNA specific, in our test conditions DNA was found to interfere in DAPK1and CCL2 assay, as confirmed by: 1) analysis of the size of the amplified PCR products on agarose gel electrophoresis; 2) running a PCR reaction with human DNA; and 3) running a PCR reaction with subjects’ samples but omitting the reverse transcription (data not shown). The DNA interference was confirmed after finishing all the experiments. Interestingly, the new lot number of reagents purchased after informing the producer about the uncertainty, no longer showed DNA interference (data not shown). Because we used all cDNA samples, it was not possible to repeat the measurements with the new lot number of reagents. On the other hand, it was not possible to study in detail the relative contribution of DNA and cDNA to the signals obtained with the old lot number of reagents, because we used all of them.
The laboratory findings are summarized in Figure 1. DAPK1 and CCL2 plasma levels of nucleic acids (mRNA and DNA) were successfully determined in all patients and in 31 (out of 33) controls. Patients had, on average, 5.1-times higher plasma levels of DAPK1 nucleic acids than controls (median and interquartile range 33.82 (11.40-77.44) vs. 6.63 (2.87-11.47); P < 0.001). Patients with diabetes had 5.5-times higher levels than patients without without diabetes (68.23 (34.12-99.18) vs. 12.38 (7.09-35.48); P = 0.006), and the latter still had 86.7 % higher levels than the controls (12.38 (7.09-35.48) vs. 6.63 (2.87-11.47); P = 0.014). On a contrary, plasma levels of CCL2 nucleic acids tended to be lower in patients than in controls (12.55 (6.77-38.61) vs. 36.25 (5.90-74.54); P = 0.062) and, when only statin-treated patients were compared to the controls, the decrease reached the significance (-66.3%; 12.21 (6.41-26.60) vs. 36.25 (5.90-74.54); P = 0.041). It is noteworthy that 22 statin-treated patients (out of 24) were receiving aspirin and 16 patients ACE inhibitors.
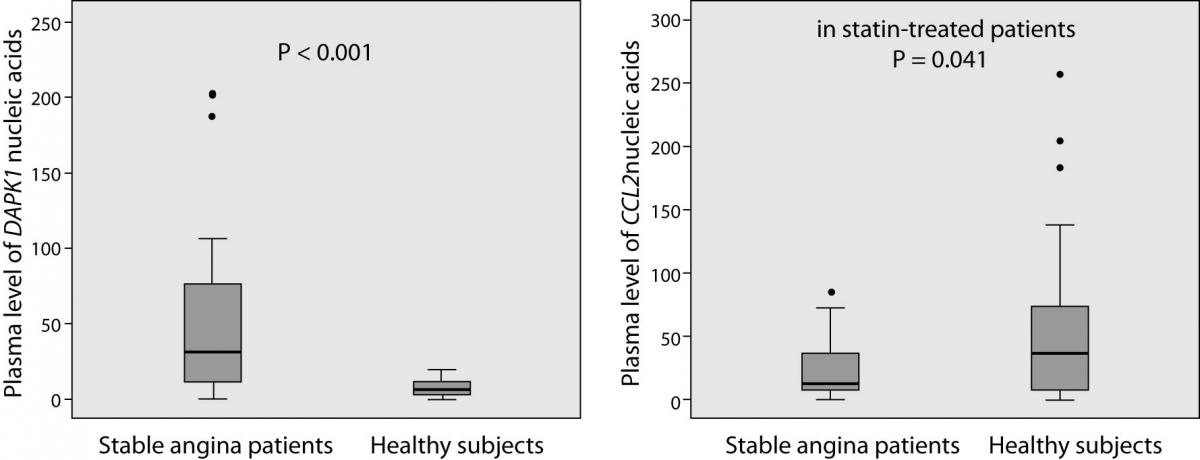
Figure 1. Plasma DAPK1 and CCL2 nucleic acids levels in patients with stable angina (N = 29) and in healthy subjects (N = 33). Plasma levels were standardized to the amount of reference gene (PPIA mRNA). Box plots demonstrate interquartile range (25th and 75th percentiles), and line inside the box denotes the median bar. The whiskers demonstrate the largest and the smallest values inside the 1.5 times interquartile range. Outliers (·) are the values between the 1.5 and the 3 times interquartile range. Extreme values are not presented.
Levels of nucleic acids in patients’ plasma did not correlate with the number of diseased coronary arteries or with any other observed parameter. In the controls, plasma levels of nucleic acids correlated neither with plasma lipids nor with any other observed parameter. Importantly, plasma levels of nucleic acids did not correlate with leukocyte or platelet counts.
Discussion
Our study shows the existence of cell-free nucleic acids from DAPK1 and CCL2, i.e. from genes responsible for atherosclerotic plaque development and destabilization, in plasma of patients with coronary atherosclerosis. Plasma nucleic acids analysis is an emerging field for non-invasive in vivo assessment and monitoring of pathologic processes in tissues of their origin. It represents an alternative or a supplement to protein analysis. Nowadays, quantitative and qualitative analysis of cell-free nucleic acids is considered to be clinically useful in non-invasive prenatal diagnosis,and in diagnosis and monitoring of numerous cancers (5), but novel applications are emerging also in other critical care settings, such as acute coronary syndrome (6–8,11). These nucleic acids can be measured precisely in only 200 mL of plasma of patients with stable angina as well as of healthy subjects (8,11). We believe that with the use of higher plasma volumes for nucleic acids isolation (commercial kits are available) and with further improvement of specificity of assays for PCR quantification there will be possible to reliably measure plasma mRNA from DAPK1 and CCL2, thus offering a new technique for non-invasive in vivo assessment and monitoring of gene expression profile in vascular beds.
Although conclusions are not as expected and DNA interference introduces uncertainty to the data, the results of our study are interesting and explicable in the light of current knowledge. High plasma levels of DAPK1 nucleic acids in patients with stable angina are consistent with the current evidence that apoptosis is a prominent feature of atherosclerotic plaque, especially in regions with a high density of macrophages (1,12). Apoptosis is an important source of nucleic acids (mRNA and DNA) in plasma (5). Furthermore, DAPK1 is a well known positive mediator of apoptosis in a wide array of apoptotic systems (13) and DABK1 was found to be highly expressed in carotid atherosclerotic plaques (2). Finally, prolonged hyperglycaemia and/or impaired insulin signalling were also shown to induce apoptosis (14). In this respect it should be noted that our study design does not exclude other sources of DAPK1 nucleic acids. DAPK1 nucleic acids containing apoptotic bodies in non-vascular tissues may escape the elimination by reticuloendothelial system in those tissues and thus protrude into the blood. This could be an additional reason for extremely high plasma DAPK1 nucleic acids levels in our patients with diabetes. Unexpectedly, stable angina patients had lower plasma levels of CCL2 nucleic acids than controls. Here it is noteworthy that if DNA had had a major contribution to the signals obtained, it would have been difficult to explain the divergent results for plasma nucleic acids levels from CCL2 and DAPK1. Lower plasma nucleic acids levels from CCL2 in patients with stable angina could be explained as a consequence of statin pleiotropy (15). Twenty-four patients (out of the 29) were receiving statin therapy. Statin- treated patients had lower plasma concentrations of CCL2 than those without statin therapy (16), and statin treatment significantly diminished neointimal and medial CCL2 in a rabbit model of atherosclerosis (17). Finally, statin treatment reduced CCL2 mRNA levels in peripheral blood monocytes in vivo (18).
To conclude, cell-free nucleic acids from genes responsible for atherosclerotic plaque development and destabilization are possible to quantify in plasma of patients with stable angina and of healthy subjects and the estimated levels are explicable in terms of current knowledge. Further studies with specific assays for PCR quantification are reasonable.
Acknowledgment
The authors acknowledge financial support from the Slovenian Research Agency (project No. L3-0129-2413-08).
Notes
Potential conflict of interest
None declared.
References
1. Kockx MM, Herman AG. Apoptosis in atherosclerosis: beneficial or detrimental? Cardiovasc Res 2000;45:736–46.
2. Martinet W, Schrijvers DM, De Meyer GR, Thielemans J, Knaapen MW, Herman AG, Kockx MM. Gene expression profiling of apoptosis-related genes in human atherosclerosis: upregulation of death-associated protein kinase. Arterioscler Thromb Vasc Biol 2002;22:2023–9.
3. Gonzalez-Quesada C, Frangogiannis NG. Monocyte chemoattractant protein-1/CCL2 as a biomarker in acute coronary syndromes. Curr Atheroscler Rep 2009;11:131–8.
4. Chen YL, Chang YJ, Jiang MJ. Monocyte chemotactic protein-1 gene and protein expression in atherogenesis of hypercholesterolemic rabbits. Atherosclerosis 1999;143:115–23.
5. Tong YK, Lo YM. Diagnostic developments involving cell-free (circulating) nucleic acids. Clin Chim Acta 2006;363:187-96.
6. Rainer TH, Lam NY, Man CY, Chiu RW, Woo KS, Lo YM. Plasma beta-globin DNA as a prognostic marker in chest pain patients. Clin Chim Acta 2006;368:110–3.
7. Wang GK, Zhu JQ, Zhang JT, Li Q, Li Y, He J, et al. Circulating microRNA: a novel potential biomarker for early diagnosis of acute myocardial infarction in humans. Eur Heart J 2010;31:659–66.
8. Stern I, Marc J, Kranjec I, Zorman D, Cerne A, Cerne D. Increased plasma levels of CATS mRNA but not CATB mRNA in patients with coronary atherosclerosis. Clin Biochem 2010;43:1427-30.
9. Cerne D, Stern I, Kranjec I, Marc J. Optimisation of methods for quantifying plasma mRNA levels from genes responsible for coronary artery plaque development and destabilization. Med Glas Ljek komore Zenicko-doboj kantona 2011;8:90-6.
10. Dovc-Drnovsek T, Emersic B, Rozman P, Cerne D, Lukac-Bajalo J. Optimization of purification of human cell-free mRNA from plasma. Ann N Y Acad Sci 2008;1137:125-9.
11. Cerne D, Stern I, Marc J, Cerne A, Zorman D, Krzisnik-Zorman S, Kranjec I. CTSS activation coexists with CD40 activation in human atheroma: Evidence from plasma mRNA analysis. Clin Biochem 2011;44:438-40.
12. Mallat Z, Tedgui A. Apoptosis in the vasculature: mechanisms and functional importance. Br J Pharmacol 2000;130:947–62.
13. Cohen O, Inbal B, Kissil JL, Raveh T, Berissi H, Spivak-Kroizaman T, et al. DAP-kinase participates in TNF-alpha- and Fas-induced apoptosis and its function requires the death domain. J Cell Biol 1999;146:141–8.
14. Hasnan J, Yusof MI, Damitri TD, Faridah AR, Adenan AS, Norbaini TH. Relationship between apoptotic markers (Bax and Bcl-2) and biochemical markers in type 2 diabetes mellitus. Singapore Med J 2010;51:50–5.
15. Kirmizis D, Chatzidimitriou D. Pleiotropic vasoprotective effects of statins: The chicken or the egg? Drug Des Devel Ther 2009;3:191–204.
16. Lewandowski M, Kornacewicz-Jach Z, Millo B, Zielonka J, Czechowska M, Kaliszczak R, et al. The influence of low dose atorvastatin on inflammatory marker levels in patients with acute coronary syndrome and its potential clinical value. Cardiol J 2008;15:357–64.
17. Bustos C, Hernández-Presa MA, Ortego M, Tuñón J, Ortega L, Pérez F, et al. HMG-CoA reductase inhibition by atorvastatin reduces neointimal inflammation in a rabbit model of atherosclerosis. J Am Coll Cardiol 1998;32:2057–64.
18. Hoogeveen RC, Morrison A, Boerwinkle E, Miles JS, Rhodes CE, Sharrett AR, Ballantyne CM. Plasma MCP-1 level and risk for peripheral arterial disease and incident coronary heart disease: Atherosclerosis Risk in Communities study. Atherosclerosis 2005;183:301–7.